How Sampling From Bioreactors Are Shifting To Rapid Automation
Processing samples to be analyzed by various offline devices during a bioprocess run is very labor intensive.
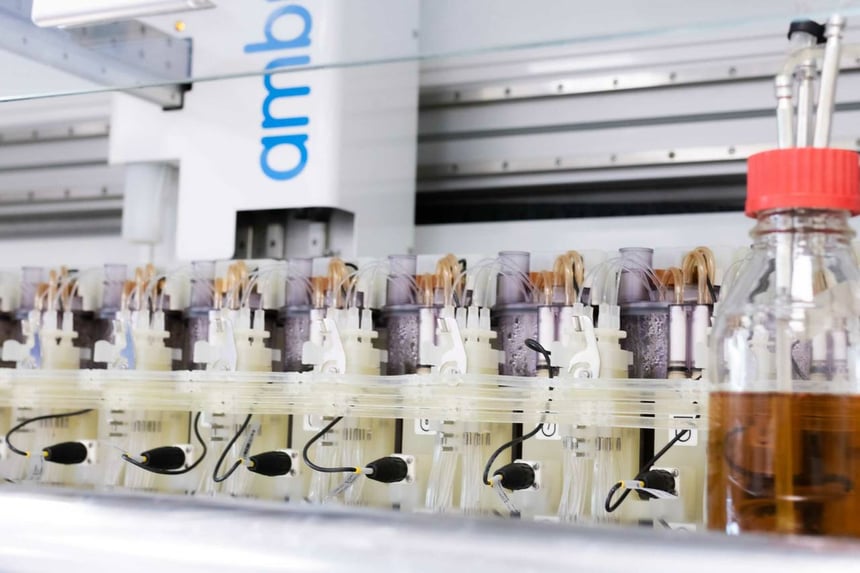
Picture yourself running a bioprocess campaign that involves a 24-way Ambr® 250 system. You set it up to collect daily samples and head off to enjoy your weekend.
Upon return to the lab on Monday you face an enormous list of 72 samples - including the sample collected that day to process on your multiple offline devices.
- Will you know which sample corresponds to which analytical experiment?
- Does everything correlate?
- Did you make any mistakes?
It’s critically important to know exactly how your process is performing. Your cells could be deteriorating or the feeding strategy you have implemented may not be enough to support the cell growth. You won’t be able to make informed decisions without this dataset at hand.
Whether you’re using single or multi-parallel bioreactor systems, all the activities incurred during this process are prone to error. Furthermore, if you’re manually preparing samples for metabolite analysis on the Cedex Bio HT or checking cell viability on the NucleoCounter®, many other steps in your process could be lost in the digital representation if not captured.
The byproduct of this process is often if not always is a very tired and frustrated scientist. But it doesn’t stop there as everyday you’ll be manually processing samples of your bioreactors until the end of that one bioprocess experiment. Now imagine that you have to run more than one experiment and repeat the same steps. There has to be a better way to manage all of this.

In order to optimize your upstream bioprocessing strategy, a fundamental knowledge of the process and its characteristics is essential. To understand that knowledge you need to first see the data and interpret it in a way that allows you to make informed decisions.
Bioreactors generate a wealth of data from multiple sensors. For example, a typical Sartorius Ambr® 250 bioreactor system will contain tens of thousands of data per bioreactor. A typical mammalian process will last 14 days or even longer so the data pool really starts to amplify the longer you run your process.
Moreover, each bioreactor is then coupled with in-/at-/off- line sensors which again contributes to the huge amount of data generated. Inline sensors are part of the bioreactors themselves such as pH probes and temperature sensors. Atline sensors are integrated devices that will analyze samples automatically sampled from your bioreactors to measure metabolites and cell counts.
An example of this device is the Nova Biomedical BioProfile® FLEX2 which can be integrated into the Sartorius Ambr® 250 bioreactor system. Offline sensors such as the Cedex Bio HT Analyzer are stand-alone units where bioreactor samples are manually supplied and the subsequent data requires intervention to link back to the rest of the bioprocessing data.
Metabolite analysis is extremely critical in process optimization as are measurements of yield, waste, cell count, and feed (usually glucose) which will indicate the health of the cells and how they are reacting to the particular setpoints in the bioprocess design.
These insights can even help predict the outcome of a process and provide feedback to the bioreactors for managing proportional integral derivative (PID) control loops. Such control loops can for example initiate feeding if glucose concentrations in a bioreactor fall below a certain threshold.

Current Solutions
Monitoring and control of critical process parameters as well as product attributes are still the basic tasks in bioprocess development. Recent trends in automation and digitalization in bioprocess have brought significant improvements by reducing human error and increasing through-put.
To solve and remove manual interventions, there are a few labs that deploy custom solutions where they attempt to adopt a streamlined approach for processing bioreactor samples.This mostly involves a liquid handling robot such as a Tecan system or Hamilton for performing high throughput wet lab experiments on the bioreactor samples, and then sending them to an analytical device for data analysis. This improves lab productivity but is limited to just a set of procedures.
Any deviation from the procedures involves weeks in developing a new protocol and testing to ensure no failures. Another downside is the lack of digitalization in which these protocols, the data and the analysis sit in localized devices and don’t utilize cloud technology and adopt FAIR principles (Findable, Accessible, Interoperable, and Reusable). Joining data across siloed instruments and data points is often a time consuming and hazardous process fraught with errors in an effort to standardize media if these principles are aren’t considered.
Process Analytical Technology (PAT) is applied in the biopharmaceutical industry for process monitoring and is essential for continuous bioprocessing and for adopting Quality by Design (QbD) approaches. The Numera from Securecell is an example of using automated sampling systems that are connected to bioreactors that automatically sample and analyze to feed back into the process. Samples are tracked and the data is unified with the bioreactor dataset.
This technology has been relatively successful in real time monitoring of bioreactor cultures. However these systems aren’t flexible when it comes down to the method of analysis. You can’t dilute, mix or move liquids around for other processes. And each unit of operation requires an additional module to be attached to the system, which means more instruments for labs to maintain.
How biopharma companies are tackling the digitalization of bioprocessing data
Processing Your Samples on a Cloud Platform
Automation and digitalization are becoming increasingly important in the modern environment of bioprocesses 4.0. Driven by the PAT initiative and the goal of achieving biologics via a QbD approach with a well-understood and optimally controlled process, the digital transformation of bioprocesses is inevitable. But this all comes at a cost which labs may not have—these systems are expensive.
But there is another way. Most labs now have liquid handling robots such as Tecans, Hamiltons or Gilsons. What if you could use sample liquid handling protocols to achieve almost real time monitoring of your bioreactor coupled with data analysis of your process?
The ideal scenario should have bioreactor samples integrate into a cloud platform and ingest into liquid handling workflows. In this manner, the platform gives users control of the liquid handlers to process multiple experiments in one single master workflow all stemming from the bioreactor sample itself. The scientist can then track where and when each bioreactor sample went.
Now that they can track the samples, scientists can adapt their single workflow to include other analytical experiments, streamlining the journey from bioreactor-to-sample-to-data and essentially achieving digitalization.
Scientists are then freed to do other tasks whilst their liquid handler automatically splits bioreactor samples into the experimental stages that they want to analyze. During or after the experiments are complete, the data can be imported into that same cloud platform for the scientist to unify and visualize over a well deserved coffee break.
Bioprocess 4.0 is on course to becoming a reality. Many are now looking to this approach to streamline their sampling upstream and downstream to maximize experiment campaigns and their own time effectively, relieving bandwidth from the seemingly never ending cascade of problems and focusing more on progressing experiments through their pipeline to conclusions. Eventually, this will become the norm in many laboratories across the world.
Want to explore this topic further? Get in touch with us and lets have an open discussion about bioprocessing.



